A comparison of deep learning u-net architectures for posterior segment oct retinal layer segmentation
A comparison of deep learning u-net architectures for posterior segment oct retinal layer segmentation"
- Select a language for the TTS:
- UK English Female
- UK English Male
- US English Female
- US English Male
- Australian Female
- Australian Male
- Language selected: (auto detect) - EN
Play all audios:
ABSTRACT Deep learning methods have enabled a fast, accurate and automated approach for retinal layer segmentation in posterior segment OCT images. Due to the success of semantic
segmentation methods adopting the U-Net, a wide range of variants and improvements have been developed and applied to OCT segmentation. Unfortunately, the relative performance of these
methods is difficult to ascertain for OCT retinal layer segmentation due to a lack of comprehensive comparative studies, and a lack of proper matching between networks in previous
comparisons, as well as the use of different OCT datasets between studies. In this paper, a detailed and unbiased comparison is performed between eight U-Net architecture variants across
four different OCT datasets from a range of different populations, ocular pathologies, acquisition parameters, instruments and segmentation tasks. The U-Net architecture variants evaluated
include some which have not been previously explored for OCT segmentation. Using the Dice coefficient to evaluate segmentation performance, minimal differences were noted between most of the
tested architectures across the four datasets. Using an extra convolutional layer per pooling block gave a small improvement in segmentation performance for all architectures across all
four datasets. This finding highlights the importance of careful architecture comparison (e.g. ensuring networks are matched using an equivalent number of layers) to obtain a true and
unbiased performance assessment of fully semantic models. Overall, this study demonstrates that the vanilla U-Net is sufficient for OCT retinal layer segmentation and that state-of-the-art
methods and other architectural changes are potentially unnecessary for this particular task, especially given the associated increased complexity and slower speed for the marginal
performance gains observed. Given the U-Net model and its variants represent one of the most commonly applied image segmentation methods, the consistent findings across several datasets here
are likely to translate to many other OCT datasets and studies. This will provide significant value by saving time and cost in experimentation and model development as well as reduced
inference time in practice by selecting simpler models. SIMILAR CONTENT BEING VIEWED BY OTHERS WIDE-FIELD OCT VOLUMETRIC SEGMENTATION USING SEMI-SUPERVISED CNN AND TRANSFORMER INTEGRATION
Article Open access 24 February 2025 SEGMENTATION OF MACULAR NEOVASCULARIZATION AND LEAKAGE IN FLUORESCEIN ANGIOGRAPHY IMAGES IN NEOVASCULAR AGE-RELATED MACULAR DEGENERATION USING DEEP
LEARNING Article Open access 01 July 2022 DEEP LEARNING SEGMENTATION OF HYPERAUTOFLUORESCENT FLECK LESIONS IN STARGARDT DISEASE Article Open access 05 October 2020 INTRODUCTION Optical
coherence tomography (OCT) is a non-invasive, high resolution imaging modality that is commonly used in ophthalmic imaging. By allowing easy visualisation of the retinal tissue, OCT scans
can be employed by clinicians and researchers for diagnosis of ocular diseases and monitoring disease progression or response to therapy through the quantitative and qualitative analysis of
features within these images. The inner and outer boundaries of the retinal and choroidal layers are of particular interest in OCT image analysis. However, the marking of the position of
these boundaries can be slow and inefficient when performed manually by a human expert annotator. Therefore, rapid and reliable automated algorithms are necessary to perform retinal
segmentation in a time-efficient manner. Early automatic methods for OCT retinal and choroidal segmentation relied upon standard image processing techniques as part of methods that were
handcrafted to suit particular sets of images1,2,3,4,5,6,7,8,9. While these methods achieve the goal of saving time, the specific nature of the rules that make up their algorithms means that
they may not generalise well to other data without manually recalibrating the algorithm, which can take considerable time and present notable difficulties. On the other hand, a range of
alternative methods have been proposed which are based on deep learning. A deep learning method can automatically learn the rules from a dataset, rectifying one of the main drawbacks of
traditional analysis techniques. Additionally, extending an algorithm to new data typically requires simply extending the training set with the new samples, while most other training and
model parameters do not need to be modified for the method to operate. There are numerous deep learning methods that have been proposed for OCT retinal segmentation with a few different
techniques commonly employed including patch-based methods10,11,12,13,14 and semantic segmentation methods14,15,16,17,18,19,20,21,22,23,24,25,26, among others. Semantic segmentation methods,
in particular, have demonstrated state-of-the-art performance in retinal and choroidal layer segmentation tasks of OCT images14. The majority of semantic segmentation methods adopt an
encoder-decoder deep neural network structure, most of which base their architectures on the U-Net27, improving over the original fully-convolutional network approach28. In the U-Net, skip
connections improve gradient flow and allow the transfer of information between the down-sampling and up-sampling paths by connecting each pair of encoder-decoder layers. The U-Net takes an
input OCT image and classifies each individual pixel in the image into one of several possible classes, thereby segmenting the image into the different regions. For OCT retinal layer
segmentation, the classes correspond to the different tissue layers and other regions around the tissue layers of interest. The output of the U-Net is a pixel-level map providing precise
segmentation of the regions and layers of interest. There are several variants which aim to improve performance compared to the original (vanilla) U-Net architecture. Dense U-Net29,30
connects all convolution block outputs to the inputs of all subsequent blocks within each level which encourage feature reuse as well as aiding in preventing vanishing gradients. The
Inception U-Net31 employs multiple convolutional kernel sizes in parallel paths at each level ensuring a greater level of robustness to images of different scales. The Attention U-Net32
employs an attention module within each skip connection, with these modules allowing for greater focus to be given to the more important spatial regions within an image and lesser focus on
those of lower importance. The residual U-Net33,34 adopts residual learning in each of the layers by adding shortcut connections which aim to improve gradient flow and work under the theory
that the residual mapping is easier to learn than the original one. The recurrent-residual (R2) U-Net35 combines the benefits of residual learning (see Residual U-Net) and recurrent
connections, the latter of which allows for improved feature accumulation and parameter utilisation. The U-Net++36 uses a set of dense skip connections allowing for improved aggregation of
features across different semantic scales. The squeeze + excite (SE) U-Net37,38,39 incorporates squeeze + excite blocks at the output of each convolutional block. These blocks allow for
activation map reweighting of both spatial locations and channels based on the relative importance of each spatial location and channel. A summary of each architecture as well as references
to their prior applications can be found in Table 1. Previous studies for medical image segmentation (including OCT) have proposed a number of variants of the U-net architecture to improve
performance. However, the effect of these U-Net architectural changes on OCT image segmentation performance is unclear, as an unbiased comparison across multiple OCT datasets has not been
performed. Indeed, previous comparisons use different datasets to one another, compare using only a single dataset, compare only using a small subset of methods, and/or contain bias due to
network architectures not being properly matched (e.g. using a different number of layers to one another). In this study, a comparison is performed using eight significant U-Net variants on
four different and varied OCT datasets to obtain an understanding of their effect on segmentation performance as well as trade-offs with respect to computational time (training and
evaluation) and complexity. The four OCT datasets encompass data from a range of different populations, ocular pathologies, scanning parameters, instruments and segmentation tasks, to ensure
that the comparison is not biased and limited to just a single dataset. The effect of the number of convolutional layers is also examined, by comparing two and three layers per block, as a
simple complementary experiment alongside the more complex architectural changes. The overall goal of this study is to determine general conclusions for semantic OCT retinal layer
segmentation using U-Net architectures which can be applied to any OCT dataset and result in significant time savings for future studies. This study also examines network architectures that
have not previously been applied to this problem. To the best of our knowledge, the U-Net++, R2U-Net and Inception U-Net have not been previously applied to OCT retinal or choroidal
segmentation and hence will be investigated in this study. METHODS DATA Four datasets were used in this study, which aim to provide a wide range of image qualities and features. Thus, it
will allow an improved understanding of the performance of the U-Net variants. For the relevant ethics information, please refer to the original studies using the citations below. All
methods were performed in accordance with the relevant guidelines and regulations. DATASET 1: HEALTHY This dataset, from a previous study49, consists of spectral domain OCT (SD-OCT) B-scans
from 104 healthy children. Approval from the Queensland University of Technology human research ethics committee was obtained before commencement of the study, and written informed consent
was provided by all participating children and their parents. All participants were treated in accordance with the tenets of the Declaration of Helsinki. Data was sourced across four
separate visits over a period of approximately 18 months, however for the purposes of this study, we utilise data from the first visit only. For this visit, 6 cross-sectional scans were
acquired, equally spaced in a radial pattern, and centred upon the fovea. Scans were acquired using the Heidelberg Spectralis SD-OCT instrument. For all scans, 30 frames were averaged using
the inbuilt automated real time function (ART) to reduce noise while enhanced depth imaging (EDI) was used to improve the visibility of the choroidal tissue. Each scan measures 1536 pixels
wide by 496 pixels deep (approximately 8.8 × 1.9 mm respectively in physical dimensions with a vertical scale of 3.9 µm per pixel and a horizontal scale of 5.7 µm per pixel). After data
collection, all scans were annotated by an expert human observer with boundary annotations created for three tissue layer boundaries including: the inner boundary of the inner limiting
membrane (ILM), the outer boundary of retinal pigment epithelium (RPE), and the choroid-sclera interface (CSI). For each scan, a semantic segmentation mask (same pixel dimensions) is
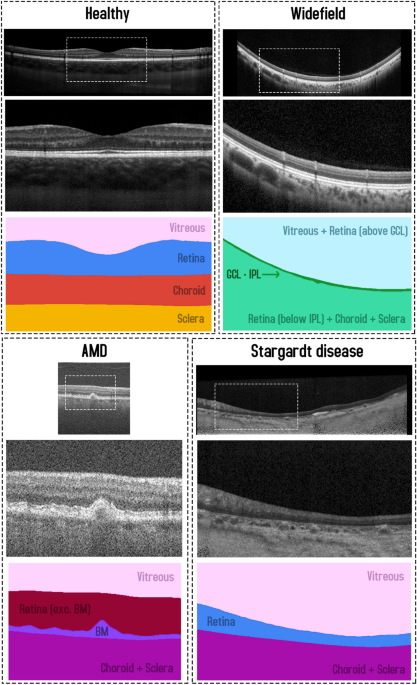
constructed with four regions: (1) vitreous (all pixels between top of the image and the ILM), (2) retina (ILM to RPE), (3) choroid (RPE to CSI), and (4) sclera (CSI to bottom of the image).
An example scan and corresponding segmentation mask is provided in Fig. 1. For the purposes of this study, the data was randomly divided into a training (40 participants, 240 scans total),
validation (10 participants, 60 scans total) and testing set (49 participants, 294 scans total) with no participant’s data overlapping between sets. DATASET 2: STARGARDT DISEASE This
dataset, described in a prior study15, consists of SD-OCT scans of patients with varying stages of Stargardt disease. Approval to identify and use SD-OCT images from patients with
genetically confirmed Stargardt disease for developing segmentation methods was obtained from the Human Ethics Office of Research Enterprise, The University of Western Australia (RA/4/1/8932
and RA/4/1/7916) and the Human Research Ethics Committee, Sir Charles Gairdner Hospital (2001-053). Scans were acquired from 18 participants, with 4 volumes (captured at different visits at
different times) of ~ 50–60 B-scans from each. The scans measure 1536 pixels wide by 496 pixels high, with a vertical scale of 3.9 µm per pixel and a horizontal scale of 5.7 µm per pixel,
this corresponds to an approximate physical area of size 8.8 × 1.9 mm (width × height). Scans were acquired using the Heidelberg Spectralis SD-OCT instrument with the ART algorithm used to
enhance the definition of each B-scan by averaging nine OCT images. Scans were taken in high-resolution mode, unless it was determined necessary to use high-speed mode owing to poor fixation
(any low-resolution scan was resized to match the resolution of the dataset). EDI was not employed. Contrast enhancement is utilised15,50 in an effort to improve layer visibility and
subsequently improve segmentation performance. After acquisition, each scan was annotated by an expert human observer, with annotations provided for two layer boundaries including: the inner
boundary of the inner limiting membrane (ILM), and the outer boundary of the retinal pigment epithelium (RPE). In some patients with Stargardt disease, the RPE and the outer retinal layers
are lost in the central region of the retina. In such cases, the remaining Bruch’s membrane (BM) that separates the residual inner retina from the choroid is marked as the outer boundary.
For each scan, a semantic segmentation mask is constructed (same pixel dimensions) with three regions: (1) vitreous (all pixels from top of image to the ILM), (2) retina (from ILM to RPE/BM)
and (3) choroid/sclera (RPE/BM to bottom of the image). An example scan and corresponding segmentation mask is provided in Fig. 1. Each participant is categorised into one of two categories
based on retinal volume (low or high). This is calculated based on the total macular retinal volume based on the boundary annotations such that there is an even number of participants in
each category. For this study, the data is divided into training (10 participants, 2426 scans), validation (2 participants, 486 scans), and testing (6 participants, 1370 scans) sets ensuring
that there is an even split of low and high volume participants in each set. DATASET 3: AGE-RELATED MACULAR DEGENERATION This dataset consists of OCT scans of patients exhibiting
age-related macular degeneration (AMD)51. All scans were acquired using the Bioptigen SD-OCT with data sourced from four different sites with scanning resolutions varying slightly between
the sites52. No image averaging is employed. A total of 269 participants are utilised, each with a single volume of 100 scans. However, only a subset of the scans is used. A scan is used
only if it contains at least 512 pixels (approximately half the width) of available boundary annotations, otherwise it is discarded. Each scan is then cropped to a central 512 pixels
horizontally with each scan also measuring 512 pixels in height (no cropping). Each scan is supplied with boundary annotations for three layer boundaries including the inner boundary of the
ILM, the outer boundary of the retinal pigment epithelium drusen complex (RPEDC) and the outer boundary of BM. For each scan, a semantic segmentation mask is constructed (same pixel
dimensions) with four regions including: (1) vitreous (all pixels from the top of the image to the ILM), (2) retina (ILM to RPEDC), (3) Bruch’s membrane (RPEDC to BM), and (4) choroid/sclera
(BM to bottom of the image). An example scan and corresponding segmentation mask is provided in Fig. 1. A total of 163 participants (4439 scans) are used for training, 54 participants (1481
scans) for validation, and 52 participants (1421 scans) for testing with all participants assigned randomly with no overlap or duplication. DATASET 4: WIDEFIELD This dataset, which has been
described in detail elsewhere53, included 12 healthy participants that had widefield OCT volume scans acquired using a widefield objective lens while maintaining central fixation. All
participants provided written consent as per protocols approved by the University of New South Wales Australia Human Research Ethics Advisory panel, and the study adhered to the tenets of
the Declaration of Helsinki. Scans were acquired using the Heidelberg Spectralis where the applied scanning protocol acquired 109 B-scans spaced 120 μm apart, spanning a total area of 55°
horizontally and 45° vertically (15.84 mm wide and 12.96 mm high), and ART was set to 16 to reduce noise. OCT B-scans were manually segmented to extract two boundaries of interest: the inner
boundary of the ganglion cell-inner layer (GCL), and the outer boundary of the inner plexiform layer (IPL). The thickness data between these two boundaries (ganglion cell-inner plexiform
layer thickness) can inform the detection and screening of a number of retinal diseases such as glaucoma54, central retinal vein occlusion55 and diabetic macular edema56. After resizing and
cropping to the region of interest, the scans measure 1392 pixels wide by 496 pixels high. For each scan, a semantic segmentation mask is constructed (same pixel dimensions) with three
regions: (1) all pixels from the top of the image to the GCL, (2) all pixels between GCL and IPL, and (3) all pixels between the IPL and the bottom of the image. An example scan and
corresponding segmentation mask is provided in Fig. 1. Scans from a total of 12 participants (~ 109 scans each, with one scan discarded from a single participant) are utilised with 6
participants assigned for training (654 images total), 3 for validation (326 images total) and 3 for testing (327 images total). There is no overlap of participants between the three sets.
TRAINING AND ARCHITECTURE PARAMETERS All networks are trained to perform semantic segmentation of the OCT scans. Using a 1 × 1 convolution layer (1 × 1 strides, filter count corresponding to
number of regions) followed by a softmax activation, the output of each network consists of class probabilities corresponding to each pixel in the original input OCT image. For the
Stargardt disease and widefield datasets, the probabilities represent the classification of each pixel into one of three areas/regions of the OCT images, while there are four such areas for
the healthy and AMD datasets as described in the Data section. For each of the eight variants of the U-net architecture that are considered, we build off previous code implementations as
follows: (1) baseline57, (2) dense58, (3) attention59, (4) squeeze and excite (SE)60, (5) residual61, (6) recurrent-residual (R2)62, (7) U-Net++63, and (8) Inception64. For the comparison in
this study, all networks utilise max pooling layers (2 × 2) for down-sampling and transposed convolutions for up-sampling (2 × 2 kernel and strides). Each network consists of four
pooling/up-sampling layers with 16 filters for all convolutions in the first layer which is doubled after each pooling and subsequently halved at each up-sampling layer. Each convolution
uses Glorot uniform65 kernel initialisation, and each layer, with the exception of the output layer, is followed by batch normalisation66 and a rectified linear unit activation. We compare
two variants for each architecture by considering two and three convolutions per layer. A visual summary of each architecture is given in Fig. 2. For the residual variant, the ‘ReLU before
addition’ block variant is selected while two recurrences are performed for the R2 variant. For the SE variant, we employ the scSE module (concurrent cSE and sSE) with default ratio of 2 for
the cSE component. The U-Net++ architecture employs the same filters within the encoder and decoder (16 filters first layer, subsequently doubled in each layer) as the other architectures.
However, all the intermediate layers (between the encoder and decoder) use 16 filters each, for computational reasons. For training, the Adam optimizer67 is employed with default parameters
(except for the learning rate which is set at 0.0001). A batch size of two is used and all networks are trained for 200 epochs, minimising cross-entropy as the training objective. For model
selection, we use the epoch with the best Dice coefficient on the validation set. The Dice coefficient is a measure of similarity between two samples (in this case, the predicted
segmentation map and ground truth map) and is defined as: $$Dice=\frac{2TP}{2TP+FP+FN}$$ (1) where TP: number of true positives, FP: number of false positives, and FN: number of false
negatives. For regularisation during training, all training samples are shuffled randomly at the beginning of each epoch. No augmentation is employed for any experiments for simplicity. The
comparison between architectures is performed by comparing the Dice coefficient on the testing set using the model selected from the best epoch. To perform a fair comparison between the
network architectures, it is necessary to run each experiment several times to ensure that there is no bias as a result of random weight initialisation and sample shuffling. Hence, all
experiments are performed three times in this study. All experiments are undertaken using Tensorflow 2.3.1 in Python 3.7.4 using an NVIDIA M40 graphics processing unit (GPU). To
statistically compare the segmentation outcomes (Dice coefficient) from the different architectures, a one-way repeated measures analysis of variance (ANOVA) was run for each of the
different datasets for both the 2 layer and 3 layer variants separately, examining the within-subject effect of network architecture. Bonferroni-adjusted pairwise comparisons were conducted
to compare between specific network architectures. An additional ANOVA was conducted to compare between the segmentation outcomes of the 2 layer and 3 layer variants. RESULTS Tables 2, 3, 4
and 5 provide a summary of the main results for the healthy dataset, Stargardt disease dataset, AMD dataset and widefield dataset, respectively. The accuracy represents the mean (and
standard deviation) overall Dice coefficient as the main performance metric, while the times (epoch training and evaluation time) together with the number of parameters allow for a
comparison of the computational complexity of the networks. Figure 3 provides a visual summary of accuracy vs. evaluation time vs. network complexity with a subplot for each of the four
datasets. It can be noted that the general clustering of the two-layer variants (squares) and three-layer variants (circles) indicates that most architectures exhibit comparable segmentation
accuracy (x-axis) with the notable exception being the R2 variant (in yellow), which showed a marginal performance improvement. Additionally, the general difference between these two
clusters (squares and circles) indicates that the three-layer variant (circles) consistently outperforms the two-layer variant (squares), but only by a small amount. While the R2 variant
(yellow) is the most accurate, it is also the slowest with respect to evaluation time (y-axis) and therefore lies in the top right corner of the subplots. However, this is not a general
trend with the Inception variant, for instance, lying in the top left corner of the subplots (slow but with relatively low accuracy compared to the other variants). Observing the subplots,
there does not appear to be any clear trends with respect to the number of network parameters (relative sizes of the circles and squares). Figures 4, 5, 6, and 7 give some example
segmentation outputs overlaid with transparency on the original OCT scans and comparisons to ground truth segmentation, demonstrating the robustness and accuracy of the segmentations for
each of the four datasets (healthy, Stargardt disease, AMD and widefield) respectively using the R2 U-Net (3 layer variant). The repeated measures ANOVA revealed a significant effect of
network architecture for both the healthy and Stargardt disease datasets (for both the 2 and 3 layer variants). Although the magnitude of differences were small, the R2 architecture was
found to be statistically significantly more accurate compared to some of the other architectures including the vanilla, attention, SE and residual U-Net architectures (all _p_ < 0.05).
There were also statistically significant differences for each architecture comparing the two and three layer variants with the three-layer variant being slightly but statistically
significantly more accurate on both the healthy and Stargardt disease datasets (_p_ < 0.05). Additionally, it was found that using three layers (compared to two layers) was significantly
more accurate overall (not taking architecture into account) (_p_ < 0.05) on the healthy, Stargardt disease and widefield datasets. On the other hand, there were no statistically
significant differences between architectures for the AMD and widefield datasets (all _p_ > 0.05). The lack of significance of the results on these datasets is likely related to the more
variable performance on the AMD dataset and the small number of participants in the widefield dataset. DISCUSSION In this study, eight different U-Net variants were compared for their
segmentation performance on four different OCT image datasets encompassing a range of ocular pathologies and scanning parameters. Each architecture was also compared using both two and three
convolutional blocks per layer. The results suggest that there is largely comparable performance (measured using Dice coefficient) for all of the architectures on the individual datasets.
Indeed, despite the increased complexity and running time of a number of the U-Net variants, their performance was not notably greater than the baseline vanilla U-Net, which already exhibits
excellent performance for OCT retinal layer segmentation. The eight U-Net architectures were similar in terms of accuracy, and the performance of each improved slightly on all four datasets
by using an additional convolutional layer per block (three instead of two), a comparatively simple architectural change. However, this was again at the cost of increased complexity and
running time as a result of the extra parameters introduced by the additional layers. Despite the resultant performance improvements for each individual network, the comparison between each
of them remains relatively unchanged. That is, their relative performance to one another is comparable as it is with two convolutional layers. The findings further highlight the importance
of comparisons between different network architectures being performed carefully (e.g. using the same number of layers). For instance, a vanilla U-Net and a Residual U-Net are often fitted
(by default) with two and three convolutional layers per block, respectively, which will potentially bias the performance of the Residual U-Net. In this specific scenario, the additional
layer is the cause of improvement as opposed to the addition of residual connections. It may be possible for additional layers (e.g. 4 or more) to be added to further improve performance,
however, this is at the cost of drastically increased model complexity as well as slower inference and training time. At a certain point, this increased complexity will also result in the
inability of the computing hardware to handle the model size (i.e. insufficient VRAM). Additionally, we hypothesise that diminishing returns are likely with respect to performance (given the
excellent performance with 3 layers) making the trade-off between performance and complexity even less favourable with higher numbers of convolutional layers. While segmentation accuracy
(Dice coefficient) is a critical metric, it is not the only one that should be considered when comparing these architectures. Indeed, training time, evaluation speed, and model complexity
are some other vital factors to consider from the perspective of real-world training and model deployment both in research and clinical practice. Small performance improvements may not be
worthwhile in certain applications if this is obtained at the cost of significantly increased complexity and reduced speed. In this case, while the vanilla U-Net is the fastest to train, one
of the fastest to evaluate, and possesses the fewest parameters (lowest complexity), its performance is still comparable and competitive with the majority of the other architectures across
all datasets. On the other hand, the R2 U-Net performs slightly better and contains only slightly more parameters but is significantly slower to train and evaluate as a result of the
recurrences where layers are reused, requiring additional computation at each step. In fact, it is the slowest to evaluate of any of the architectures highlighting a key trade-off when
selecting this architecture. Overall, the Inception U-Net does not perform favourably in any metric possessing significantly more parameters than most of the other networks, being the
slowest to train and one of the slowest to evaluate, and yielding comparable performance on all of the datasets. Similarly, although the Dense U-Net uses more parameters than most of the
other networks (as it takes in more feature maps in the subsequent dense block layers), its performance is only comparable across all four datasets. As expected, there is a high correlation
between training and evaluation time but there are a few minor exceptions. Although the Inception U-Net is the slowest to train, the R2 U-Net (3 layer variant) is the slowest to evaluate
across all four datasets. This is likely due to differences in the type of operations that are performed in the forward pass (used in evaluation) vs. in backpropagation (not used in
evaluation). Based on these findings there is clear a trade-off between memory, training time, evaluation speed, and performance. However, the improvements in performance are small across
all tested datasets and models, suggesting that simpler, faster models are the optimal choices for OCT retinal layer segmentation. The general level of performance between the datasets
varies. However, this is expected given the higher difficulty and subjectivity associated with images exhibiting pathology and with poorer image quality (i.e. varying noise levels). The
comparison between architectures on some datasets told a slightly different story in some cases. For instance, we observe a small benefit to using the squeeze + excitation U-Net on the
Stargardt disease dataset, similar to a previous study15. It appears that the recalibration of feature maps is beneficial in the presence of highly pathological data and where the overall
segmentation accuracies (Dice coefficients) are notably lower than the other datasets. However, the overall level of improvement is small. In general, the performance between the three runs
for each experiment was largely consistent, with standard deviations of less than 0.1% on three of the four datasets with respect to the Dice coefficient. However, the performance on the AMD
dataset shows greater variability, particularly for the case of two convolutional layers on some of the architectures. Indeed, the vanilla, residual, attention and inception variants all
exhibited standard deviations > 0.2% when outfitted with two convolutional layers. Adding a third convolutional layer showed a marked reduction in variability on all architectures (except
Inception, where the number of layers does not apply). We hypothesise that this variability is caused by the greater noise present in the AMD dataset, with this problem rectified
significantly with the increased network learning capacity (i.e. by adding an extra layer). We note that there are also differences in the training and inference speeds for each of the
different datasets which are associated with differing quantities of data and different sized images respectively. However, these differences are irrelevant for an architectural comparison
which examines a single dataset at a time, and has no effect on the overall conclusions of this study. In this study, four datasets were employed that are highly representative of real world
OCT datasets that may be encountered in practice. These are also similar to those used in other studies involving deep learning based OCT segmentation methods11,12,14,15. Some of these
datasets contain only a few participants, indicative of the common difficulty of sourcing large numbers of participants due to privacy and confidentiality reasons as well as a lack of
readily available patients exhibiting less common ocular pathologies. However, a significant number of scans from each of these patients are used, to support invariance of the model to
patient agnostic image features and artefacts, such as speckle noise. Despite potential issues surrounding a low number of participants and model bias, the performance of all models is
excellent across all datasets suggesting that this does not degrade the performance. Additionally, the findings from the architectural comparison on each dataset are largely comparable,
demonstrating that the lower quantity of data on some datasets does not appear to be biasing the results or invalidating the findings of this study. This also further supports the previously
demonstrated fact that the U-Net can be trained well even with a relatively small number of images27. Here, there was an exclusive focus on an architectural comparison between U-Net
variants for OCT semantic segmentation of retinal layers. We stress that there are other OCT segmentation methods that have not been considered here. These specific methods are not
considered as they incorporate other changes and modifications beyond the architecture (e.g. loss function, class weighting, additional post-processing etc.) which are outside the scope of
this study. For instance, another study68 has performed a comparison between traditional (Dice and logistic) and adversarial (GAN) losses. Future studies should examine other such parameters
and perform similar comparisons. There are also numerous other configurations of U-Net architectures (e.g. different residual block configurations, number of recurrences) and other semantic
segmentation architectures (e.g. DeepLabv3+69, SegNet70) that have not been tested here to retain a manageable scope for this study. An interesting direction for future work could be to
examine an ensemble approach using different U-Net architectures for each ensemble member. However, given the largely comparable performance between the tested architectures in this study,
we speculate that any performance benefits may be minor. CONCLUSION In this study, we have performed a comprehensive, unbiased comparison of U-Net architecture variants for the application
of semantic segmentation in retinal OCT images across several datasets. All tested U-Net architectures provide excellent performance for OCT retinal layer segmentation, and the results
suggest that there is little notable difference in performance between them when evaluated using the Dice metric. There are expected trade-offs between performance, speed and complexity that
are important to consider depending on the particular clinical and research application as well as constraints on time and available hardware. The findings of this study also highlight the
importance of careful and unbiased comparisons of deep learning methods and correctly matching network architectures to obtain a true understanding on the impact of network architectural
changes. Overall, the significantly increased complexity and reduced speed of the U-Net variants with only marginal performance gains suggest that the baseline vanilla U-Net is an optimal
choice for OCT retinal layer segmentation in practice. State-of-the-art models do not appear to provide a clear benefit for this application while increasing the number of layers in each
model resulted in small performance gains but, again, with a trade-off with respect to complexity and speed. A significant time investment in any study involving deep learning is that of
model comparison and optimal selection. The findings in this study can help to provide a solid, well-informed starting point, alleviating the time and cost burden of experimental comparison
in future studies while also guiding model selection towards simpler and faster models. The findings here are largely consistent across several varied OCT datasets with differing
pathologies, instruments, scanning parameters and segmentation tasks. Therefore, these can be generalised and are likely transferable to a wide range of other OCT datasets further
highlighting the benefit of this study for future work with U-Net based models for OCT retinal layer segmentation which are commonly employed for this application. DATA AVAILABILITY The
healthy, Stargardt disease and widefield datasets analysed during the current study are currently not publicly available. However, the algorithms and code used throughout this study are
publicly and readily available at https://github.com/jakugel/unet-variants. REFERENCES * Koozekanani, D., Boyer, K. & Roberts, C. Retinal thickness measurements from optical coherence
tomography using a Markov boundary model. _IEEE Trans. Med. Imaging_ 20, 900–916 (2001). Article CAS PubMed Google Scholar * Oliveira, J., Pereira, S., Gonçalves, L., Ferreira, M. &
Silva, C. A. Multi-surface segmentation of OCT images with AMD using sparse high order potentials. _Biomed. Opt. Express_ 8, 281–297 (2017). Article PubMed Google Scholar * Cabrera
Fernández, D., Salinas, H. M. & Puliafito, C. A. Automated detection of retinal layer structures on optical coherence tomography images. _Opt. Express_ 13, 10200–10216 (2005). Article
ADS PubMed Google Scholar * Kafieh, R., Rabbani, H., Abramoff, M. D. & Sonka, M. Intra-retinal layer segmentation of 3D optical coherence tomography using coarse grained diffusion
map. _Med. Image Anal._ 17, 907–928 (2013). Article PubMed Google Scholar * Chiu, S. J. _et al._ Automatic segmentation of seven retinal layers in SDOCT images congruent with expert
manual segmentation”. _Opt. Express_ 18, 19413–19428 (2010). Article ADS PubMed PubMed Central Google Scholar * Niu, S., de Sisternes, L., Chen, Q., Leng, T. & Rubin, D. L.
Automated geographic atrophy segmentation for SD-OCT images using region-based C-V model via local similarity factor”. _Biomed. Opt. Express_ 7, 581–600 (2016). Article PubMed PubMed
Central Google Scholar * Chiu, S. J. _et al._ Kernel regression based segmentation of optical coherence tomography images with diabetic macular edema. _Biomed. Opt. Express_ 6, 1172–1194
(2015). Article PubMed PubMed Central Google Scholar * Li, K., Wu, X., Chen, D. Z. & Sonka, M. Optimal surface segmentation in volumetric Images—A graph theoretic approach. _IEEE
Trans. Pattern Anal. Mach. Intell._ 28, 119–134 (2006). Article CAS PubMed PubMed Central Google Scholar * Tian, J. _et al._ Real-time automatic segmentation of optical coherence
tomography volume data of the macular region. _PLoS ONE_ 10, e0133908 (2015). Article PubMed PubMed Central Google Scholar * Fang, L. _et al._ Automatic segmentation of nine retinal
layer boundaries in OCT images of non-exudative AMD patients using deep learning and graph search. _Biomed. Opt. Express_ 8, 2732–2744 (2017). Article PubMed PubMed Central Google Scholar
* Hamwood, J., Alonso-Caneiro, D., Read, S. A., Vincent, S. J. & Collins, M. J. Effect of patch size and network architecture on a convolutional neural network approach for automatic
segmentation of OCT retinal layers. _Biomed. Opt. Express_ 9, 3049–3066 (2018). Article PubMed PubMed Central Google Scholar * Kugelman, J., Alonso-Caneiro, D., Read, S. A., Vincent, S.
J. & Collins, M. J. Automatic segmentation of OCT retinal boundaries using recurrent neural networks and graph search. _Biomed. Opt. Express_ 9, 5759–5777 (2018). Article PubMed PubMed
Central Google Scholar * Masood, S. _et al._ Automatic choroid layer segmentation from optical coherence tomography images using deep learning. _Sci. Rep._ 9, 1–18 (2019). Google Scholar
* Kugelman, J. _et al._ Automatic choroidal segmentation in OCT images using supervised deep learning methods. _Sci. Rep._ 9, 13298 (2019). Article ADS PubMed PubMed Central Google
Scholar * Kugelman, J. _et al._ Retinal boundary segmentation in Stargardt disease optical coherence tomography images using automated deep learning. _Trans. Vis. Sci. Tech._ 9, 12 (2020).
Article Google Scholar * Devalla, S. K. _et al._ DRUNET: a dilated residual U-Net deep learning network to segment optic nerve head tissues in optical coherence tomography images. _Biomed.
Opt. Express_ 9, 3244–3265 (2018). Article PubMed PubMed Central Google Scholar * Venhuizen, F. G. _et al._ Robust total retina thickness segmentation in optical coherence tomography
images using convolutional neural networks. _Biomed. Opt. Express_ 8, 3292–3316 (2017). Article PubMed PubMed Central Google Scholar * Roy, A. G. _et al._ ReLayNet: Retinal layer and
fluid segmentation of macular optical coherence tomography using fully convolutional network. _Biomed. Opt. Express_ 8, 3627–3642 (2017). Article PubMed PubMed Central Google Scholar *
Borkovkina, S., Camino, A., Janpongsri, W., Sarunic, M. V. & Jian, J. Real-time retinal layer segmentation of OCT volumes with GPU accelerated inferencing using a compressed, low-latency
neural network. _Biomed. Opt. Express_ 11, 3968–3984 (2020). Article CAS PubMed PubMed Central Google Scholar * Pekala, M. _et al._ Deep learning based retinal OCT segmentation.
_Comput. Biol. Med_ 114, 103445 (2019). Article CAS PubMed Google Scholar * Sousa, J. A. _et al._ Automatic segmentation of retinal layers in OCT images with intermediate age-related
macular degeneration using U-Net and DexiNed. _PLoS ONE_ 16, e0251591 (2021). Article CAS PubMed PubMed Central Google Scholar * Apostolopoulos, S., De Zanet, S., Ciller, C., Wolf, S.
& Sznitman, R. Pathological OCT retinal layer segmentation using branch residual U-shape networks. https://arxiv.org/1707.04931 (2017). * Alsaih, K., Yusoff, M. Z., Tang, T. B., Faye, I.
& Mériaudeau, F. Deep learning architectures analysis for age-related macular degeneration segmentation on optical coherence tomography scans. _Comput. Methods Programs Biomed._ 195,
105566 (2020). Article CAS PubMed Google Scholar * Zuehua, W., Xiangcong, X., Yaguang, Z., Dingan, H. A new method with SEU-Net model for automatic segmentation of retinal layers in
optical coherence tomography images in _Proceedings of the 2021 IEEE 2__nd__ Internation Conference on Big Data, Artifical Intelligence and Internet of Things Engineering (ICBAIE)_, 260–263
(IEEE, 2021). * Mishra, Z. _et al._ Automated retinal layer segmentation using graph-based algorithm incorporating deep-learning-derived information. _Sci. Rep._ 10, 9541 (2020). Article
ADS PubMed PubMed Central Google Scholar * Chen, M. _et al._ Multiscale dual attention mechanism for fluid segmentation of optical coherence tomography images. _Appl. Opt._ 60, 6761–6768
(2021). Article ADS PubMed Google Scholar * Ronneberger, O., Fischer, P. & Brox, T. U-Net: convolutional networks for biomedical image segmentation.https://arxiv.org/abs/1505.04597
(2015). * Shelhamer, E., Long, J. & Darrell, T. Fully convolutional networks for semantic segmentation. _IEEE Trans. Pattern Anal. Mach. Intell._ 39, 640–651 (2016). Article PubMed
Google Scholar * Huang, G., Liu, Z., van der Maaten, L. & Weinberger, K. Q. Densely connected convolutional networks. https://arxiv.org/abs/1608.06993 (2016). * Cai, S. _et al._
Dense-UNet: a novel multiphoton in vivo cellular image segmentation model based on a convolutional neural network. _Quant. Imaging Med. Surg._ 10, 1275–1285 (2020). Article PubMed PubMed
Central Google Scholar * Szegedy, C. _et al._ Going deeper with convolutions. https://arxiv.org/abs/1409.4842 (2014). * Oktay, O. _et al._ Attention U-Net: Learning where to look for the
pancreas. https://arxiv.org/abs/1804.03999 (2018). * He, K., Zhang, X., Ren, S. & Sun, J. Deep residual learning for image recognition in _2016 IEEE Conference on Computer Vision and
Pattern Recognition,_ 770–778 (IEEE, 2016). * He, K., Zhang, X., Ren, S. & Sun, J. Identity mappings in deep residual networks in _2016 European Conference on Computer Vision_. 630–645
(Springer, 2016). * Alom, M. Z., Hasan, M., Yakopcic, C., Taha, T. M. & Asari, V. K. Recurrent residual convolutional neural network based on U-Net (R2U-Net) for medical image
segmentation. https://arxiv.org/abs/1802.06955 (2018). * Zhou, Z., Siddiquee, M. M. R, Tajbakhsh, N. & Liang, J. UNet++: A nested U-Net architecture for medical image segmentation.
https://arxiv.org/abs/1807.10165 (2018). * Hu, J., Shen, L. & Sun, G. Squeeze-and-excitation networks. https://arxiv.org/abs/1709.01507 (2017). * Roy, A.G., Navab, N. & Wachinger, C.
Concurrent spatial and channel squeeze & excitation in fully convolutional networks, in _2018 International Conference on Medical Image Computing and Computer-Assisted Intervention
(MICCAI)_, 421–429 (Springer, 2018). * Roy, A. G., Navab, N. & Wachinger, C. Recalibrating fully convolutional networks with spatial and channel ‘squeeze & excitation’
blocks.https://arxiv.org/abs/1808.08127 (2018). * Guan, S., Khan, A. A., Sikdar, S. & Chitnis, P. V. Fully dense UNet for 2D sparse photoacoustic tomography artefact removal.
https://arxiv.org/abs/1808.10848 (2018). * Cao, Y., Liu, S., Peng, Y. & Li, J. DenseUNet: densely connected Unet for electron microscopy image segmentation. _IET Image Process._ 14,
2682–2689 (2020). Article Google Scholar * Punn, N. S. & Agarwal, S. Inception U-Net architecture for semantic segmentation to identify nuclei in microscopy cell images. _ACM Trans.
Multim. Comput. Commun. Appl._ 16, 1–15 (2020). Article Google Scholar * Delibasoglu, I. & Cetin, M. Improved U-Nets with inception blocks for building detection. _J. Appl. Remote
Sens._ 14, 044512 (2020). Article ADS Google Scholar * Guo, C. _et al._ SA-UNet: spatial attention U-net for retinal vessel segmentation. https://arxiv.org/abs/2004.03696 (2020). * Hu,
J., Song, Y., Zhang, L., Bai, S. & Yi, Z. Multi-scale attention U-net for segmenting clinical target volume in graves’ ophthalmopathy. _Neurocomputing_ 427, 74–83 (2021). Article Google
Scholar * Wang, C. & Gan, M. Tissue self-attention network for the segmentation of optical coherence tomography images on the esophagus. _Biomed. Opt. Express_ 12, 2631–2646 (2021).
Article PubMed PubMed Central Google Scholar * Zhang, Z., Liu, Q. & Wang, Y. Road extraction by deep residual U-Net. https://arxiv.org/abs/1711.10684 (2017). * Alom, M. Z., Hasan,
M., Yakopcic, C., Taha, T. M. & Asari, V. K. Nuclei segmentation with recurrent residual convolutional neural networks based U-Net (R2U-Net) in _Proceedings of the IEEE National
Conference on Aerospace and Electronics (NAECON)._ 228–233 (IEEE, 2018). * Read, S. A., Collins, M. J., Vincent, S. J. & Alonso-Caneiro, D. Choroidal thickness in childhood. _Invest.
Ophthalmol. Vis. Sci._ 54, 3586–3593 (2013). Article PubMed Google Scholar * Girard, M. J., Strouthidis, N. G., Ethier, C. R. & Mari, J. M. Shadow removal and contrast enhancement in
optical coherence tomography images of the human optic nerve head. _Invest. Ophthalmol. Vis. Sci._ 52, 7738–7748 (2011). Article PubMed Google Scholar * Farsiu, S. _et al._ Quantitative
classification of eyes with and without intermediate age-related macular degeneration using optical coherence tomography. _Ophthalmology_ 121, 162–172 (2014). Article PubMed Google Scholar
* Chiu, S. J. _et al._ Validated automatic segmentation of AMD pathology including drusen and geographic atrophy in SD-OCT images. _Invest. Opthalmol. Vis. Sci._ 53, 53–61 (2012). Article
Google Scholar * Tong, J., Yoshioka, N., Alonso-Caneiro, D. & Zangerl, B. Ganglion cell-inner plexiform layer measurements derived from widefield compared to montaged 9-field optical
coherence tomography. _Clin. Exp. Optom._ https://doi.org/10.1080/08164622.2021.1993058 (2021). Article PubMed Google Scholar * Yang, Z. _et al._ Diagnostic ability of macular ganglion
cell inner plexiform layer measurements in glaucoma using swept source and spectral domain optical coherence tomography. _PLoS ONE_ 10, e0125957 (2015). Article PubMed PubMed Central
Google Scholar * Chan, E. W. _et al._ Disorganization of retinal inner layers and ellipsoid zone disruption predict visual outcomes in central retinal vein occlusion. _Ophthalmol. Retina._
3, 83–92 (2019). Article PubMed Google Scholar * Das, R., Spence, G., Hogg, R. E., Stevenson, M. & Chakravarthy, U. Disorganisation of inner retina and outer retinal morphology in
diabetic macular edema. _JAMA Ophthalmol._ 136, 202–208 (2018). Article PubMed PubMed Central Google Scholar * zhixuhao, Implementation of deep learning framework —Unet, using Keras,
https://github.com/zhixuhao/unet (2019). * clguo, Fully Dense UNet implementation in medical image segmentation, https://github.com/clguo/Dense_Unet_Keras (2019). * Xu, X. Attention U-Net.
https://www.kaggle.com/xxc025/attention-u-net (2019). * titu1994, Implementation of Squeeze and Excitation Networks in Keras, https://github.com/titu1994/keras-squeeze-excite-network (2020).
* S. Taghipour, UNet_ResNet34, https://www.kaggle.com/saeedtqp/unet-resnet34 (2019). * lixiaolei1982, Keras Implementation of U-Net, R2U-Net, Attention U-Net, Attention R2U-Net,
https://github.com/lixiaolei1982/Keras-Implementation-of-U-Net-R2U-Net-Attention-U-Net-Attention-R2U-Net (2019). * longpollehn, A Keras implementation of UNet++,
https://github.com/longmakesstuff/UnetPlusPlus (2021). * danielenricocahall, Keras-UNet, https://github.com/danielenricocahall/Keras-UNet (2019). * Glorot, X. & Bengio, Y. Understanding
the difficulty of training deep feedforward neural networks in _Proceedings of the thirteenth international conference on artificial intelligence and statistics._ 249–256 (JMLR, 2010). *
Ioffe, S. & Szegedy, C. Batch normalization: Accelerating deep network training by reducing internal covariate shift. https://arxiv.org/abs/1502.03167 (2015). * Kingma, D. P. & Ba,
J. Adam: A method for stochastic optimization. https://arxiv.org/abs/1505.00393 (2014). * Kugelman, J., Alonso-Caneiro, D., Read, S. A., Vincent, S. J. & Collins, M. J. OCT
chorio-retinal segmentation using adversarial loss in _2021 Digital Image Computing: Techniques and Applications (DICTA)_. * Chen, L. –C., Zhu, Y., Papandreou, G., Schroff, F. & Adam, H.
Encoder-decoder with atrous separable convolution for semantic image segmentation. https://arxiv.org/abs/1802.02611 (2018). * Badrinarayanan, V., Kendall, A. & Cipolla, R. SegNet: a
deep convolutional encoder-decoder architecture for image segmentation. https://arxiv.org/abs/1511.00561 (2015). Download references ACKNOWLEDGEMENTS Computational resources and services
used in this work were provided in part by the HPC and Research Support Group, Queensland University of Technology, Brisbane, Australia. We gratefully acknowledge support from the NVIDIA
Corporation for the donation of GPUs used in this work. This work was supported by Ideas Grants and a Fellowship from the Australian National Health and Medical Research Council of Australia
[GNT1186915, GNT1116360 (FC) and MRF1142962 (FC)], a scholarship from the Australian Government Research Training Program (JT), and project grants from the Telethon-Perth Children’s
Hospital Research Fund (DAC, FC), and the Macular Disease Foundation Australia (FC). Additionally, Guide Dogs NSW/ACT provided a PhD scholarship (JT), salary support (JP and MK) and support
clinical service delivery at the Centre for Eye Health. The funding bodies had no role in the conceptualization or writing of the paper. AUTHOR INFORMATION AUTHORS AND AFFILIATIONS * Contact
Lens and Visual Optics Laboratory, Centre for Vision and Eye Research, School of Optometry and Vision Science, Queensland University of Technology (QUT), Kelvin Grove, Australia Jason
Kugelman, Joseph Allman, Scott A. Read, Stephen J. Vincent, Michael J. Collins & David Alonso-Caneiro * Centre for Eye Health, University of New South Wales (UNSW), Sydney, NSW,
Australia Janelle Tong & Michael Kalloniatis * School of Optometry and Vision Science, UNSW, Sydney, NSW, Australia Janelle Tong & Michael Kalloniatis * Centre for Ophthalmology and
Visual Science (Incorporating Lions Eye Institute), The University of Western Australia, Perth, WA, Australia Fred K. Chen & David Alonso-Caneiro * Department of Ophthalmology, Royal
Perth Hospital, Perth, WA, Australia Fred K. Chen * Ophthalmology, Department of Surgery, University of Melbourne, East Melbourne, Victoria, Australia Fred K. Chen Authors * Jason Kugelman
View author publications You can also search for this author inPubMed Google Scholar * Joseph Allman View author publications You can also search for this author inPubMed Google Scholar *
Scott A. Read View author publications You can also search for this author inPubMed Google Scholar * Stephen J. Vincent View author publications You can also search for this author inPubMed
Google Scholar * Janelle Tong View author publications You can also search for this author inPubMed Google Scholar * Michael Kalloniatis View author publications You can also search for this
author inPubMed Google Scholar * Fred K. Chen View author publications You can also search for this author inPubMed Google Scholar * Michael J. Collins View author publications You can also
search for this author inPubMed Google Scholar * David Alonso-Caneiro View author publications You can also search for this author inPubMed Google Scholar CONTRIBUTIONS Design of the
research: D.A.C., J.K. Software development: J.A., J.K. Analysis and interpretation of data: J.K., D.A.C., S.A.R., J.A. Drafting the manuscript: J.K., D.A.C. Reviewed and approved final
manuscript: J.K., J.A., S.A.R., S.J.V, J.T., M.K., F.K.C., M.J.C., D.A.C. CORRESPONDING AUTHOR Correspondence to Jason Kugelman. ETHICS DECLARATIONS COMPETING INTERESTS The authors declare
no competing interests. ADDITIONAL INFORMATION PUBLISHER'S NOTE Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
RIGHTS AND PERMISSIONS OPEN ACCESS This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and
reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes
were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material.
If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to
obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/. Reprints and permissions ABOUT THIS ARTICLE CITE THIS
ARTICLE Kugelman, J., Allman, J., Read, S.A. _et al._ A comparison of deep learning U-Net architectures for posterior segment OCT retinal layer segmentation. _Sci Rep_ 12, 14888 (2022).
https://doi.org/10.1038/s41598-022-18646-2 Download citation * Received: 31 March 2022 * Accepted: 17 August 2022 * Published: 01 September 2022 * DOI:
https://doi.org/10.1038/s41598-022-18646-2 SHARE THIS ARTICLE Anyone you share the following link with will be able to read this content: Get shareable link Sorry, a shareable link is not
currently available for this article. Copy to clipboard Provided by the Springer Nature SharedIt content-sharing initiative
Trending News
Tom schwartz reveals whether he’d marry again after katie maloney divorceEXPLORE MORE Tom Schwartz may be off the market — forever. The “Vanderpump Rules” star confessed to Andy Cohen on “Watch...
Public sector finances, uk: february 2025 time seriesAccredited official statistics PUBLIC SECTOR FINANCES, UK: FEBRUARY 2025 TIME SERIES The time series datasets consistent...
Ballot Flaws Reported - Los Angeles TimesThe City Council directed the city attorney to investigate after City Treasurer Alice DeLong complained of format incons...
San francisco keeps e-cigarette ban as voters reject repeal measure once backed by juulSan Francisco’s upcoming ban on the sale of e-cigarettes will move forward after voters overwhelmingly rejected a ballot...
404 - Page Not Found WInext स्पेशल और पढ़ें > horoscope4 hours ago Aaj Ka Rashifal 14 June 2025: वृष राशि वालों की आर्थिक स्थिति बेहतर होगी, श...
Latests News
A comparison of deep learning u-net architectures for posterior segment oct retinal layer segmentationABSTRACT Deep learning methods have enabled a fast, accurate and automated approach for retinal layer segmentation in po...
A laminated microfluidic device for comprehensive preclinical testing in the drug adme processABSTRACT New techniques are urgently needed to replace conventional long and costly pre-clinical testing in the new drug...
Why ranbir skipped anurag’s pujaThere have been reports that the couple has been avoiding each other at public events to Ranbir Kapoor and Katrina Kaif ...
Bradley signs ordinance to protect historic zoneMayor Tom Bradley signed an ordinance Thursday to protect the historic architectural nature of a residential neighborhoo...
Confusion over uc transfers continuesThe state reversed itself last week and said it would give 5,800 recent high school graduates the option of heading dire...